Anti-Histone H3 (phospho S28) antibody (ab5169)
Key features and details
- Rabbit polyclonal to Histone H3 (phospho S28)
- Suitable for: WB, PepArr, ICC/IF
- Reacts with: Human, Recombinant fragment
- Isotype: IgG
Overview
-
Product name
Anti-Histone H3 (phospho S28) antibody
See all Histone H3 primary antibodies -
Description
Rabbit polyclonal to Histone H3 (phospho S28) -
Host species
Rabbit -
Specificity
This antibody is specific for Histone H3 phosphorylated at residue Ser 28 and does not recognise the unmodified residue or another phosphorylated residue (Ser 10) on the same histone. -
Tested applications
Suitable for: WB, PepArr, ICC/IFmore details -
Species reactivity
Reacts with: Human, Recombinant fragment
Predicted to work with: Drosophila melanogaster
-
Immunogen
Synthetic peptide. This information is proprietary to Abcam and/or its suppliers.
-
General notes
The Life Science industry has been in the grips of a reproducibility crisis for a number of years. Abcam is leading the way in addressing this with our range of recombinant monoclonal antibodies and knockout edited cell lines for gold-standard validation. Please check that this product meets your needs before purchasing.
If you have any questions, special requirements or concerns, please send us an inquiry and/or contact our Support team ahead of purchase. Recommended alternatives for this product can be found below, along with publications, customer reviews and Q&As
Properties
-
Form
Liquid -
Storage instructions
Shipped at 4°C. Store at +4°C short term (1-2 weeks). Upon delivery aliquot. Store at -20°C or -80°C. Avoid freeze / thaw cycle. -
Storage buffer
pH: 7.40
Preservative: 0.02% Sodium azide
Constituent: PBS
Batches of this product that have a concentration < 1mg/ml may have BSA added as a stabilising agent. If you would like information about the formulation of a specific lot, please contact our scientific support team who will be happy to help. -
 Concentration information loading...
Concentration information loading... -
Purity
Immunogen affinity purified -
Clonality
Polyclonal -
Isotype
IgG -
Research areas
Associated products
-
Compatible Secondaries
-
Isotype control
-
Recombinant Protein
Applications
The Abpromise guarantee
Our Abpromise guarantee covers the use of ab5169 in the following tested applications.
The application notes include recommended starting dilutions; optimal dilutions/concentrations should be determined by the end user.
| Application | Abreviews | Notes |
|---|---|---|
| WB | (2) |
Use a concentration of 1 µg/ml. Detects a band of approximately 17 kDa (predicted molecular weight: 15 kDa).
|
| PepArr |
Use a concentration of 0.02 - 0.002 µg/ml.
|
|
| ICC/IF |
1/5000.
|
| Notes |
|---|
|
WB
Use a concentration of 1 µg/ml. Detects a band of approximately 17 kDa (predicted molecular weight: 15 kDa). |
|
PepArr
Use a concentration of 0.02 - 0.002 µg/ml. |
|
ICC/IF
1/5000. |
Target
-
Function
Core component of nucleosome. Nucleosomes wrap and compact DNA into chromatin, limiting DNA accessibility to the cellular machineries which require DNA as a template. Histones thereby play a central role in transcription regulation, DNA repair, DNA replication and chromosomal stability. DNA accessibility is regulated via a complex set of post-translational modifications of histones, also called histone code, and nucleosome remodeling. -
Sequence similarities
Belongs to the histone H3 family. -
Developmental stage
Expressed during S phase, then expression strongly decreases as cell division slows down during the process of differentiation. -
Post-translational
modificationsAcetylation is generally linked to gene activation. Acetylation on Lys-10 (H3K9ac) impairs methylation at Arg-9 (H3R8me2s). Acetylation on Lys-19 (H3K18ac) and Lys-24 (H3K24ac) favors methylation at Arg-18 (H3R17me).
Citrullination at Arg-9 (H3R8ci) and/or Arg-18 (H3R17ci) by PADI4 impairs methylation and represses transcription.
Asymmetric dimethylation at Arg-18 (H3R17me2a) by CARM1 is linked to gene activation. Symmetric dimethylation at Arg-9 (H3R8me2s) by PRMT5 is linked to gene repression. Asymmetric dimethylation at Arg-3 (H3R2me2a) by PRMT6 is linked to gene repression and is mutually exclusive with H3 Lys-5 methylation (H3K4me2 and H3K4me3). H3R2me2a is present at the 3' of genes regardless of their transcription state and is enriched on inactive promoters, while it is absent on active promoters.
Methylation at Lys-5 (H3K4me), Lys-37 (H3K36me) and Lys-80 (H3K79me) are linked to gene activation. Methylation at Lys-5 (H3K4me) facilitates subsequent acetylation of H3 and H4. Methylation at Lys-80 (H3K79me) is associated with DNA double-strand break (DSB) responses and is a specific target for TP53BP1. Methylation at Lys-10 (H3K9me) and Lys-28 (H3K27me) are linked to gene repression. Methylation at Lys-10 (H3K9me) is a specific target for HP1 proteins (CBX1, CBX3 and CBX5) and prevents subsequent phosphorylation at Ser-11 (H3S10ph) and acetylation of H3 and H4. Methylation at Lys-5 (H3K4me) and Lys-80 (H3K79me) require preliminary monoubiquitination of H2B at 'Lys-120'. Methylation at Lys-10 (H3K9me) and Lys-28 (H3K27me) are enriched in inactive X chromosome chromatin.
Phosphorylated at Thr-4 (H3T3ph) by GSG2/haspin during prophase and dephosphorylated during anaphase. Phosphorylation at Ser-11 (H3S10ph) by AURKB is crucial for chromosome condensation and cell-cycle progression during mitosis and meiosis. In addition phosphorylation at Ser-11 (H3S10ph) by RPS6KA4 and RPS6KA5 is important during interphase because it enables the transcription of genes following external stimulation, like mitogens, stress, growth factors or UV irradiation and result in the activation of genes, such as c-fos and c-jun. Phosphorylation at Ser-11 (H3S10ph), which is linked to gene activation, prevents methylation at Lys-10 (H3K9me) but facilitates acetylation of H3 and H4. Phosphorylation at Ser-11 (H3S10ph) by AURKB mediates the dissociation of HP1 proteins (CBX1, CBX3 and CBX5) from heterochromatin. Phosphorylation at Ser-11 (H3S10ph) is also an essential regulatory mechanism for neoplastic cell transformation. Phosphorylated at Ser-29 (H3S28ph) by MLTK isoform 1, RPS6KA5 or AURKB during mitosis or upon ultraviolet B irradiation. Phosphorylation at Thr-7 (H3T6ph) by PRKCBB is a specific tag for epigenetic transcriptional activation that prevents demethylation of Lys-5 (H3K4me) by LSD1/KDM1A. At centromeres, specifically phosphorylated at Thr-12 (H3T11ph) from prophase to early anaphase, by DAPK3 and PKN1. Phosphorylation at Thr-12 (H3T11ph) by PKN1 is a specific tag for epigenetic transcriptional activation that promotes demethylation of Lys-10 (H3K9me) by KDM4C/JMJD2C. Phosphorylation at Tyr-42 (H3Y41ph) by JAK2 promotes exclusion of CBX5 (HP1 alpha) from chromatin.
Monoubiquitinated by RAG1 in lymphoid cells, monoubiquitination is required for V(D)J recombination (By similarity). Ubiquitinated by the CUL4-DDB-RBX1 complex in response to ultraviolet irradiation. This may weaken the interaction between histones and DNA and facilitate DNA accessibility to repair proteins. -
Cellular localization
Nucleus. Chromosome. - Information by UniProt
-
Database links
- Entrez Gene: 8350 Human
- Entrez Gene: 8351 Human
- Entrez Gene: 8352 Human
- Entrez Gene: 8353 Human
- Entrez Gene: 8354 Human
- Entrez Gene: 8355 Human
- Entrez Gene: 8356 Human
- Entrez Gene: 8357 Human
see all -
Alternative names
- H3 histone family member E pseudogene antibody
- H3 histone family, member A antibody
- H3/A antibody
see all
Images
-
Anti-Histone H3 (phospho S28) antibody (ab5169) at 1 µg/ml + Hela Whole Cell Lysate - Colcemid Treated at 2.5 µg
Secondary
Goat Anti-Rabbit IgG H&L (HRP) (ab97051) at 1/50000 dilution
Developed using the ECL technique.
Performed under reducing conditions.
Predicted band size: 15 kDa
Observed band size: 17 kDa why is the actual band size different from the predicted?
Exposure time: 30 secondsThis blot was produced using a 4-12% Bis-tris gel under the MES buffer system. The gel was run at 200V for 35 minutes before being transferred onto a Nitrocellulose membrane at 30V for 70 minutes. The membrane was then blocked for an hour using 2% Bovine Serum Albumin before being incubated with ab5169 overnight at 4°C. Antibody binding was detected using an anti-rabbit antibody conjugated to HRP, and visualised using ECL development solution ab133406.
-
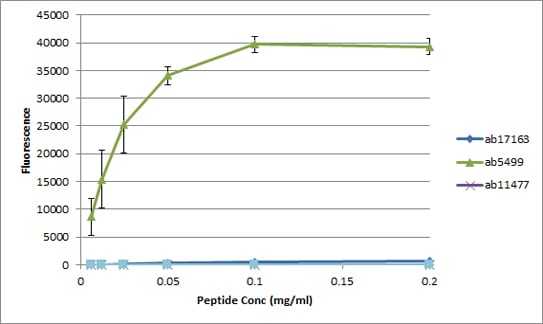
All batches of ab5169 are tested in Peptide Array against peptides to different Histone H3 modifications. Six dilutions of each peptide are printed on to the Peptide Array in triplicate and results are averaged before being plotted on to a graph. Results show strong binding to Histone H3 - phospho S28 peptide (ab5499), indicating that this antibody specifically recognises the Histone H3 - phospho S28 modification.
ab17163 - Histone H3 unmodified
ab5499 - Histone H3 - phospho S28
ab11477 - Histone H3 - phospho S10
-
 Immunocytochemistry/ Immunofluorescence - Anti-Histone H3 (phospho S28) antibody (ab5169)The image was submitted as part of a review by Krik McManus, University of British Columbia
Immunocytochemistry/ Immunofluorescence - Anti-Histone H3 (phospho S28) antibody (ab5169)The image was submitted as part of a review by Krik McManus, University of British ColumbiaIndian Muntjac (top panel) and HeLa cells (bottom panel) immunofluorescently labelled with ab5169 (green) at a working dilution of 1/5000. The DNA is counterstained with DAPI and is shown in blue in the top panel and red in the bottom panel. This antibody gives a characteristic staining pattern for Histone H3 (phospho S28) whereby the signal increases in intensity during late G2 and continues to increase until metaphase. Upon entry into anaphase the signal begins to decrease until reaching basal levels by early G1. 100x magnification.
-
 Western blot - Anti-Histone H3 (phospho S28) antibody (ab5169)Image courtesy of Richelle Sopko, Harvard University, U.S.A.All lanes : Anti-Histone H3 (phospho S28) antibody (ab5169) at 1/1000 dilution
Western blot - Anti-Histone H3 (phospho S28) antibody (ab5169)Image courtesy of Richelle Sopko, Harvard University, U.S.A.All lanes : Anti-Histone H3 (phospho S28) antibody (ab5169) at 1/1000 dilution
Lane 1 : Wild type 0-4 hour old fruit fly embryos.
Lane 2 : 0-4 hour old fruit fly embryos expressing wee RNAi
Secondary
All lanes : Donkey anti-rabbit IgG, Horseradish Peroxidase-L at 1/10000 dilution
Developed using the ECL technique.
Performed under reducing conditions.
Predicted band size: 15 kDa
Exposure time: 10 minutes0-4hr old wee shRNA embryos (lane 2) should display elevated phH3Ser28 levels relative to 0-4hr old EGFP shRNA embryos (lane 1). Blocked with 10% BSA.
-
 Immunocytochemistry/ Immunofluorescence - Anti-Histone H3 (phospho S28) antibody (ab5169)This image was submitted as part of a review by Kirk McManus, University of British Columbia
Immunocytochemistry/ Immunofluorescence - Anti-Histone H3 (phospho S28) antibody (ab5169)This image was submitted as part of a review by Kirk McManus, University of British ColumbiaIn situ peptide competition was perfomed on paraformaldehyde-fixed HeLa cells. Four 25µl aliquots were made, to which 7.5µg (1.5µl) of no peptide, H3 unmodified peptide (ab2623), H3 phospho S10 peptide (ab11477) or H3 phospho S28 peptide (ab5499) was added, mixed by vortexing and incubated for 1 hour at room temperature. Cells on glass coverslips were fixed with 4% paraformaldehyde (10min) and gently washed twice with PBS, then permeabilized with 0.5% Triton X-100 in PBS (10min) and gently washed three times with PBS. The cells were immunofluorescently labeled with either the peptide-competed antibody or the control antibody (i.e. no peptide) for 30min at room temperature, washed briefly with PBS containing 0.1% Triton X-100 (1 min) and twice with PBS. The cells were then incubated with an appropriate dilution of a secondary antibody at room temperature for 30min, rinsed as above and mounted using a 90% glycerol in PBS mount
Protocols
Datasheets and documents
-
SDS download
-
Datasheet download
References (16)
ab5169 has been referenced in 16 publications.
- Paul AS et al. Co-option of Plasmodium falciparum PP1 for egress from host erythrocytes. Nat Commun 11:3532 (2020). PubMed: 32669539
- Dong W et al. Streptococcus pneumoniae Infection Promotes Histone H3 Dephosphorylation by Modulating Host PP1 Phosphatase. Cell Rep 30:4016-4026.e4 (2020). PubMed: 32209465
- Ganter M et al. Plasmodium falciparum CRK4 directs continuous rounds of DNA replication during schizogony. Nat Microbiol 2:17017 (2017). PubMed: 28211852
- Shandilya J et al. Regulation of AURORA B function by mitotic checkpoint protein MAD2. Cell Cycle 15:2196-2201 (2016). PubMed: 27341405
- Bhattacharya S et al. Brief Communication: Featured Article: Histone H2A mono-ubiquitination and cellular transformation are inversely related in N-nitrosodiethylamine-induced hepatocellular carcinoma. Exp Biol Med (Maywood) 241:1739-44 (2016). PubMed: 27190257