Anti-KMT2A / MLL antibody (ab272023)
Key features and details
- Rabbit polyclonal to KMT2A / MLL
- Suitable for: WB, IP, ChIP/Chip
- Reacts with: Human
- Isotype: IgG
Overview
-
Product name
Anti-KMT2A / MLL antibody
See all KMT2A / MLL primary antibodies -
Description
Rabbit polyclonal to KMT2A / MLL -
Host species
Rabbit -
Tested applications
Suitable for: WB, IP, ChIP/Chipmore details -
Species reactivity
Reacts with: Human
Predicted to work with: Mouse
-
Immunogen
Synthetic peptide within Human KMT2A/ MLL aa 2725-2775. The exact sequence is proprietary. The epitope is found in the C-terminal 180 kDa fragment generated by proteolytic cleavage. The epitope is found in isoform 14P-18B of KMT2A/ MLL.
Database link: Q03164 -
Positive control
- WB: HeLa, Jurkat and HEK-293T whole cell lysate. IP: HEK-293T whole cell lysate. ChIP on ChIP: Chromatin from K562 cells.
-
General notes
The Life Science industry has been in the grips of a reproducibility crisis for a number of years. Abcam is leading the way in addressing this with our range of recombinant monoclonal antibodies and knockout edited cell lines for gold-standard validation. Please check that this product meets your needs before purchasing.
If you have any questions, special requirements or concerns, please send us an inquiry and/or contact our Support team ahead of purchase. Recommended alternatives for this product can be found below, along with publications, customer reviews and Q&As
Properties
-
Form
Liquid -
Storage instructions
Shipped at 4°C. Store at +4°C short term (1-2 weeks). Upon delivery aliquot. Store at -20°C long term. Avoid freeze / thaw cycle. -
Storage buffer
pH: 7
Preservative: 0.09% Sodium azide
Constituent: Tris citrate/phosphate
pH 7 to 8 -
 Concentration information loading...
Concentration information loading... -
Purity
Immunogen affinity purified -
Clonality
Polyclonal -
Isotype
IgG -
Research areas
Associated products
-
Compatible Secondaries
-
Positive Controls
Applications
The Abpromise guarantee
Our Abpromise guarantee covers the use of ab272023 in the following tested applications.
The application notes include recommended starting dilutions; optimal dilutions/concentrations should be determined by the end user.
| Application | Abreviews | Notes |
|---|---|---|
| WB |
1/2000 - 1/10000. Predicted molecular weight: 432 kDa.
|
|
| IP |
Use at 2-5 µg/mg of lysate.
|
|
| ChIP/Chip |
Use at an assay dependent concentration.
Use 10 µg. |
| Notes |
|---|
|
WB
1/2000 - 1/10000. Predicted molecular weight: 432 kDa. |
|
IP
Use at 2-5 µg/mg of lysate. |
|
ChIP/Chip
Use at an assay dependent concentration. Use 10 µg. |
Target
-
Function
Histone methyltransferase that plays an essential role in early development and hematopoiesis. Catalytic subunit of the MLL1/MLL complex, a multiprotein complex that mediates both methylation of 'Lys-4' of histone H3 (H3K4me) complex and acetylation of 'Lys-16' of histone H4 (H4K16ac). In the MLL1/MLL complex, it specifically mediates H3K4me, a specific tag for epigenetic transcriptional activation. Has weak methyltransferase activity by itself, and requires other component of the MLL1/MLL complex to obtain full methyltransferase activity. Has no activity toward histone H3 phosphorylated on 'Thr-3', less activity toward H3 dimethylated on 'Arg-8' or 'Lys-9', while it has higher activity toward H3 acetylated on 'Lys-9'. Required for transcriptional activation of HOXA9. Promotes PPP1R15A-induced apoptosis. -
Tissue specificity
Heart, lung, brain and T- and B-lymphocytes. -
Involvement in disease
Note=Chromosomal aberrations involving MLL are a cause of acute leukemias. Translocation t(1;11)(q21;q23) with MLLT11/AF1Q; translocation t(3;11)(p21;q23) with NCKIPSD/AF3p21; translocation t(3,11)(q25,q23) with GMPS; translocation t(4;11)(q21;q23) with AFF1/MLLT2/AF4; insertion ins(5;11)(q31;q13q23) with AFF4/AF5Q31; translocation t(5;11)(q12;q23) with AF5-alpha/CENPK; translocation t(6;11)(q27;q23) with MLLT4/AF6; translocation t(9;11)(p22;q23) with MLLT3/AF9; translocation t(10;11)(p11.2;q23) with ABI1; translocation t(10;11)(p12;q23) with MLLT10/AF10; t(11;15)(q23;q14) with CASC5 and ZFYVE19; translocation t(11;17)(q23;q21) with MLLT6/AF17; translocation t(11;19)(q23;p13.3) with ELL; translocation t(11;19)(q23;p13.3) with MLLT1/ENL; translocation t(11;19)(q23;p23) with GAS7; translocation t(X;11)(q13;q23) with FOXO4/AFX1. Translocation t(3;11)(q28;q23) with LPP. Translocation t(10;11)(q22;q23) with TET1. Translocation t(9;11)(q34;q23) with DAB2IP. Translocation t(4;11)(p12;q23) with FRYL. Fusion proteins MLL-MLLT1, MLL-MLLT3 and MLL-ELL interact with PPP1R15A and, on the contrary to unfused MLL, inhibit PPP1R15A-induced apoptosis.
Note=A chromosomal aberration involving MLL may be a cause of chronic neutrophilic leukemia. Translocation t(4;11)(q21;q23) with SEPT11. -
Sequence similarities
Belongs to the histone-lysine methyltransferase family. TRX/MLL subfamily.
Contains 3 A.T hook DNA-binding domains.
Contains 1 bromo domain.
Contains 1 CXXC-type zinc finger.
Contains 1 FY-rich C-terminal domain.
Contains 1 FY-rich N-terminal domain.
Contains 3 PHD-type zinc fingers.
Contains 1 post-SET domain.
Contains 1 SET domain. -
Domain
the 9aaTAD motif is a transactivation domain present in a large number of yeast and animal transcription factors.
The SET domain structure is atypical and is not in an optimal position to have methyltransferase activity. It requires other components of the MLL1/MLL complex, such as ASH2L or RBBP5, to order the active site and obtain optimal histone methyltransferase activity.
The CXXC-type zinc finger binds bind to nonmethyl-CpG dinucleotides. -
Post-translational
modificationsProteolytic cleavage by TASP1 generates MLL cleavage product N320 and MLL cleavage product C180, which reassemble through a non-covalent association. 2 cleavage sites exist, cleavage site 1 (CS1) and cleavage site 2 (CS2), to generate MLL cleavage products N320 and C180. CS2 is the major site. -
Cellular localization
Nucleus and Nucleus. Localizes to a diffuse nuclear pattern when not associated with MLL cleavage product N320. - Information by UniProt
-
Database links
- Entrez Gene: 4297 Human
- Entrez Gene: 214162 Mouse
- Omim: 159555 Human
- SwissProt: Q03164 Human
- SwissProt: P55200 Mouse
- Unigene: 258855 Human
- Unigene: 2389 Mouse
-
Alternative names
- ALL-1 antibody
- ALL1 antibody
- C-terminal cleavage product of 180 kDa antibody
see all
Images
-
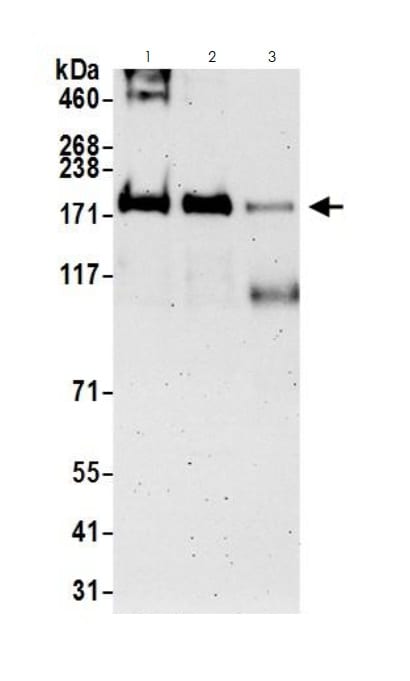
KMT2A was immunoprecipitated from HEK293T whole cell lysate (1 mg per IP reaction, 20% loaded) with ab272023 at 3 µg per reaction. Western blot was performed on the immunoprecipitate using ab272023 at 1 µg/mL.
Lane 1: rabbit anti-MLL1 IP in HEK293T whole cell lysate.
Lane 2: ab272023 IP in HEK293T whole cell lysate.
Lane 3: Control IgG in HEK293T whole cell lysate.
Detection: Chemiluminescence with an exposure time of 3 minutes
-
All lanes : Anti-KMT2A / MLL antibody (ab272023) at 0.1 µg/ml
Lane 1 : HeLa (Human epithelial cell line from cervix adenocarcinoma) whole cell lysate
Lane 2 : HEK-293T (Human epithelial cell line from embryonic kidney transformed with large T antigen) whole cell lysate
Lane 3 : Jurkat (Human T cell leukemia cell line from peripheral blood) whole cell lysate
Lysates/proteins at 50 µg per lane.
Predicted band size: 432 kDa
Exposure time: 3 minutes -
ChIP-chip scatter plot of ab272023 enriched DNA binding sites versus input reference DNA.
Plot A. 10 μg of ab272023 was used to immunoprecipitate chromatin from K562 cells according to Ren et al (Genes Dev. 2002 16: 245-256). Immunoprecipitatesd DNA and reference DNA were amplified via ligation-mediated PCR and the products labeled with fluorescent dUTPs. The labeled ChIP and reference DNA were pooled, hybridized to a DNA microarray, and analyzed. Data points below the +3 SD curve (red line) represent significantly enriched binding sites.
Plot B. As a control, a similar experiment was performed using normal rabbit IgG. Compared to the ab272023 ChIP, normal rabbit IgG showed little enrichment.
Protocols
To our knowledge, customised protocols are not required for this product. Please try the standard protocols listed below and let us know how you get on.
Datasheets and documents
-
SDS download
-
Datasheet download
References (1)
ab272023 has been referenced in 1 publication.
- Feng Z et al. Mixed Lineage Leukemia 1 Promoted Neuron Apoptosis in Ischemic Penumbra via Regulating ASK-1/TNF-a Complex. Front Neuroanat 14:36 (2020). PubMed: 32792914