NFkB p50 Transcription Factor Assay Kit (Colorimetric) (ab207217)
Key features and details
- Assay type: Semi-quantitative
- Detection method: Colorimetric
- Platform: Microplate reader
- Assay time: 3 hr 30 min
- Sample type: Cell Lysate, Nuclear Extracts
- Sensitivity: 500 ng/well
Overview
-
Product name
NFkB p50 Transcription Factor Assay Kit (Colorimetric)
See all NFkB p105 / p50 kits -
Detection method
Colorimetric -
Sample type
Cell Lysate, Nuclear Extracts -
Assay type
Semi-quantitative -
Sensitivity
< 500 ng/well -
Assay time
3h 30m -
Species reactivity
Reacts with: Mouse, Human -
Product overview
NFkB p50 Transcription Factor Assay Kit (Colorimetric) (ab207217) is a high throughput assay to quantify NFkB p50 activation in nuclear extracts. This assay combines a quick ELISA format with a sensitive and specific non-radioactive assay for transcription factor activation.
A specific double stranded DNA sequence containing the NFkB p50 consensus binding site (5´ - GGGACTTTCC - 3´) has been immobilized onto a 96-well plate. Active NFkB p50 present in nuclear or whole cell extracts specifically binds to the oligonucleotide. NFkB p50 is detected by a primary antibody that recognizes an epitope of NFkB p50 accessible only when the protein is activated and bound to its target DNA. An HRP-conjugated secondary antibody provides sensitive colorimetric readout at OD 450 nm. This product detects human and mouse NFkB p50.
Key performance and benefits:
- Assay time: 3.5 hours (cell extracts preparation not included).
- Detection limit: < 0.5 µg nuclear extract/well.
- Detection range: 0.2 – 10 µg nuclear extract/well.
-
Notes
The transcription factor NFkB is implicated in the regulation of many genes that code for mediators of the immune, acute phase and inflammatory responses. The DNA-binding protein complex recognizes a discrete nucleotide sequence (5´ - GGGACTTTCC - 3´) in the upstream region of a variety of cellular and viral response genes. NFkB is composed of homo- and heterodimeric complexes of members of the Rel (NFkB) family. There are five subunits of the NFkB family in mammals: p50, p65 (RelA), c-Rel, p52 and RelB. These proteins share a conserved 300 amino acid sequence in the N-terminal region, known as the Rel homology domain, that mediates DNA binding, protein dimerization and nuclear localization. This domain is also a target of the IkB inhibitors, which include IkBα, IkBβ, IkBγ, Bcl-3, p105 and p100.
Various dimer combinations of the NFkB subunits have distinct DNA binding specificities and may serve to activate specific sets of genes such as adhesion molecules, immunoreceptors and cytokines. The p50/p65 (NFkB1/RelA) heterodimers and the p50 homodimers are the most common dimers found in the NFkB signaling pathway. In the majority of cells, NFkB exists in an inactive form in the cytoplasm, bound to the inhibitory IkB proteins. Treatment of cells with various inducers results in the phosphorylation, ubiquitination and subsequent degradation of IkB proteins. Proteolytic cleavage of p105 results in two proteins: p50, which has DNA-binding activity but no transactivation domain, and its antagonist, the inhibitory IkBγ protein. This results in the release of NFkB dimers, which subsequently translocate to the nucleus, where they activate appropriate target genes. NFkB can be activated by a number of stimuli, including components of bacterial cell walls, such as lipopolysaccharide, or inflammatory cytokines, such as TNF-α or IL-1β.
-
Platform
Microplate reader
Properties
-
Storage instructions
Please refer to protocols. -
Components 1 x 96 tests 5 x 96 tests 10X Antibody Binding Buffer 1 x 2.2ml 1 x 11ml 10X Wash Buffer 1 x 22ml 1 x 110ml 96-well NFkB assay plate 1 unit 5 units Anti-rabbit HRP-conjugated IgG 1 x 11µl 1 x 55µl Binding Buffer 1 x 10ml 1 x 50ml Developing Solution 1 x 11ml 1 x 55ml Dithiothreitol (DTT) (1 M) 1 x 100µl 1 x 500µl Herring sperm DNA 1 x 100µl 1 x 500µl Lysis Buffer 1 x 10ml 1 x 50ml Mutated oligonucleotide (10 pmol/µL) 1 x 100µl 1 x 500µl NFkB p50 antibodies 1 x 11µl 1 x 55µl Plate sealer 1 unit 5 units Positive control nuclear extract 1 x 40µl 1 x 200µl Protease Inhibitor Cocktail 1 x 100µl 1 x 500µl Stop Solution 1 x 11ml 1 x 55ml Wild-type oligonucleotide (10 pmol/µL) 1 x 100µl 1 x 500µl -
Research areas
-
Function
NF-kappa-B is a pleiotropic transcription factor which is present in almost all cell types and is involved in many biological processed such as inflammation, immunity, differentiation, cell growth, tumorigenesis and apoptosis. NF-kappa-B is a homo- or heterodimeric complex formed by the Rel-like domain-containing proteins RELA/p65, RELB, NFKB1/p105, NFKB1/p50, REL and NFKB2/p52 and the heterodimeric p65-p50 complex appears to be most abundant one. The dimers bind at kappa-B sites in the DNA of their target genes and the individual dimers have distinct preferences for different kappa-B sites that they can bind with distinguishable affinity and specificity. Different dimer combinations act as transcriptional activators or repressors, respectively. NF-kappa-B is controlled by various mechanisms of post-translational modification and subcellular compartmentalization as well as by interactions with other cofactors or corepressors. NF-kappa-B complexes are held in the cytoplasm in an inactive state complexed with members of the NF-kappa-B inhibitor (I-kappa-B) family. In a conventional activation pathway, I-kappa-B is phosphorylated by I-kappa-B kinases (IKKs) in response to different activators, subsequently degraded thus liberating the active NF-kappa-B complex which translocates to the nucleus. NF-kappa-B heterodimeric p65-p50 and RelB-p50 complexes are transcriptional activators. The NF-kappa-B p50-p50 homodimer is a transcriptional repressor, but can act as a transcriptional activator when associated with BCL3. NFKB1 appears to have dual functions such as cytoplasmic retention of attached NF-kappa-B proteins by p105 and generation of p50 by a cotranslational processing. The proteasome-mediated process ensures the production of both p50 and p105 and preserves their independent function, although processing of NFKB1/p105 also appears to occur post-translationally. p50 binds to the kappa-B consensus sequence 5'-GGRNNYYCC-3', located in the enhancer region of genes involved in immune response and acute phase reactions. In a complex with MAP3K8, NFKB1/p105 represses MAP3K8-induced MAPK signaling; active MAP3K8 is released by proteasome-dependent degradation of NFKB1/p105. -
Sequence similarities
Contains 7 ANK repeats.
Contains 1 death domain.
Contains 1 RHD (Rel-like) domain. -
Domain
The C-terminus of p105 might be involved in cytoplasmic retention, inhibition of DNA-binding, and transcription activation.
Glycine-rich region (GRR) appears to be a critical element in the generation of p50. -
Post-translational
modificationsWhile translation occurs, the particular unfolded structure after the GRR repeat promotes the generation of p50 making it an acceptable substrate for the proteasome. This process is known as cotranslational processing. The processed form is active and the unprocessed form acts as an inhibitor (I kappa B-like), being able to form cytosolic complexes with NF-kappa B, trapping it in the cytoplasm. Complete folding of the region downstream of the GRR repeat precludes processing.
Phosphorylation at 'Ser-903' and 'Ser-907' primes p105 for proteolytic processing in response to TNF-alpha stimulation. Phosphorylation at 'Ser-927' and 'Ser-932' are required for BTRC/BTRCP-mediated proteolysis.
Polyubiquitination seems to allow p105 processing.
S-nitrosylation of Cys-61 affects DNA binding. -
Cellular localization
Nucleus. Cytoplasm. Nuclear, but also found in the cytoplasm in an inactive form complexed to an inhibitor. - Information by UniProt
-
Alternative names
- DKFZp686C01211
- DNA binding factor KBF1
- DNA binding factor KBF1 EBP1
see all -
Database links
- Entrez Gene: 4790 Human
- Entrez Gene: 18033 Mouse
- Omim: 164011 Human
- SwissProt: P19838 Human
- SwissProt: P25799 Mouse
- Unigene: 618430 Human
- Unigene: 256765 Mouse
Images
-
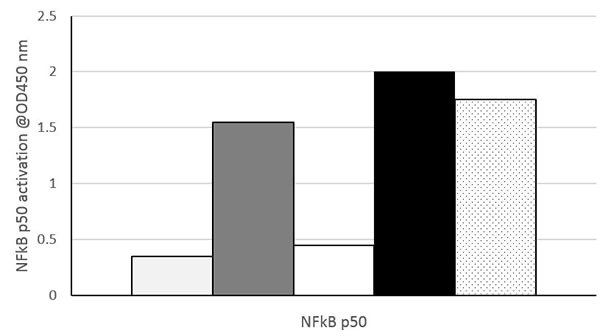
Nuclear extracts prepared from untreated HeLa cells (Light gray), HeLa cells treated with TNF-α (Dark gray), untreated Jurkat cells (White), Jurkat cells treated with PMA and calcium ionophore (CI) (Black) and Raji cells (Black dots on white) were assayed at 10 µg/well for NFkB p50 activity using ab207217. Data shown are the results from wells assayed in duplicate. These results are provided for demonstration only.
Datasheets and documents
-
SDS download
-
Datasheet download
References (3)
ab207217 has been referenced in 3 publications.
- Li Y et al. The lncARSR/PTEN/Akt/nuclear factor-kappa B feedback regulatory loop contributes to doxorubicin resistance in hepatocellular carcinoma. J Biochem Mol Toxicol 36:e23119 (2022). PubMed: 35678308
- Liu J et al. Long noncoding RNA LINC01578 drives colon cancer metastasis through a positive feedback loop with the NF-?B/YY1 axis. Mol Oncol 14:3211-3233 (2020). PubMed: 33040438
- Kumar V et al. Leishmania donovani Activates Hypoxia Inducible Factor-1a and miR-210 for Survival in Macrophages by Downregulation of NF-?B Mediated Pro-inflammatory Immune Response. Front Microbiol 9:385 (2018). PubMed: 29568285